Molecular biology is a hot topic at the moment and yet, it’s one that the general public have a hard time nailing down. It is a vast topic that encompasses a number of wider scientific areas and provides unique insights for scientists and clinicians engaged in diagnosis and treatment of various diseases. So what is it and how does it help?
Broadly speaking molecular biology as the name suggests, is a field of science in which we closely examine the molecules of life and their interactions within our bodies. The vast majority of this is done by looking at the nucleic acids that make up our genome, and the RNA and proteins which are transcribed and translated from the genome. If we can understand the underlying mechanisms of how a protein is coded for in the body, then we can almost certainly identify problems in that process when they arise. Or as happens in some cases, when a protein ceases to be produced and we expect it to be there!
The Core Techniques
Molecular biology uses a lot of really interesting techniques. A lot of molecular work is very advanced in terms of the technology, knowledge, and skill sets used in it. So what sort of techniques can we use to investigate the genome and its products? I’m going to talk about two of the most commonly used and important ones briefly. Now in all honesty, each of these techniques are complex and could easily take up numerous posts themselves but i’ll just gloss over them briefly for now.
Polymerase Chain reaction (PCR)
So we get a sample into the laboratory and regardless of what the final outcome of our investigation is, lets say we need to isolate a certain gene of interest. There can be various reasons why we need to look at this, and lots of clues it can give us but I shall discuss these separately below. For now, just know that that is the plan!
As you will already know, your genome lives nicely packaged up within the nucleus of our cells and so we need to get it out of the cells so we can work with it. When we extract DNA from samples what we get is the full three billion base pairs of DNA. Most of this is unnecessary information to us. As stated earlier we want to look at just one gene. The problem we run into, is that we simply don’t have enough sample to look at. Sure, we have our whole genome but we only need a tiny proportion of it and on top of that, we can do essentially nothing with a single copy of a gene. The answer, is to amplify that gene and make millions of copies of it. Why not make plenty of what we’re looking for eh?
That’s where PCR comes in. It is a technique used to make multiple copies of a region of interest within the genome and its based simply on the idea that we can artificially multiply genes using cycles of heating and cooling. There are three stages of a PCR reaction and we do each of them in order, at a different temperature and we repeat the cycling multiple times to give us many copies of our target sequence or gene. The diagram below, explains it very well.
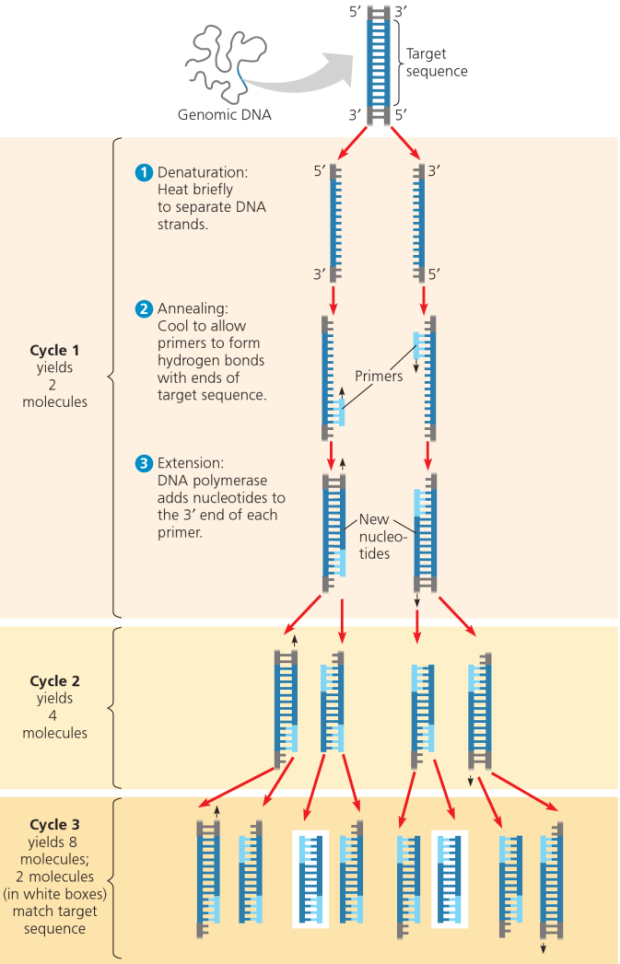
Next Generation Sequencing (NGS)
So we have plenty of copies of our target gene to work with now, but what are we actually going to do with it? This is where the next stage of the process comes into play. We need to be able to read the gene. There are various methods for doing this and at this stage I won’t be going into each of them. One of the more advanced and commonly used techniques is called Next Generation Sequencing or NGS.
As a rough insight into why this technique has been adapted by so many molecular laboratories in recent years, it is important to look at the predecessor that we had prior to implementing NGS. The previous technique was sanger seqeuncing, and it was used to sequence the entire human genome for the first time ever. A massive endeavour that was conducted by a multinational conglomerate of thirteen countries, the project took over ten years to complete and cost close to $3 billion dollars to complete (nearly a dollar a nucleotide!). With the implementation of NGS technology, we can now sequence a full human genome in around 24 hours. A major improvement!
So back to our own experiment. With our millions of copies of the gene we want to look at, we can clean up our sample during what we call “preparing the library” and then pop it onto an NGS analyser of some description and it will essentially sequence the nucleotides within our region of interest and tell us what they are.
Now I said that with great ease but I must stress the fact that library preps, although dependant on the experiment taking place, can take three or four days to complete properly. It’s an extremely important part of the experiment because all the NGS analyser will do is read the nucleotides of what we put in. If we haven’t properly cleaned up our sample and prepped our libraries i.e. made sure what is in the sample is only our region of interest, chances are we will get other readings in our results. What I would also like to stress at this point is that NGS is probably the most advanced molecular biology technique we have and i’m really doing a massive disservice to its complexity and capability by skirting over the topic with such a lack of depth. There is far too much to cover in a single post and it could be a full series of its own! if there is interest then perhaps I will cover it. But i digress.
The general idea with NGS is that when we put our sample into the analyser, it builds up the strands of DNA one nucleotide at a time and reads which nucleotide has been attached each time. When we input our sample, millions of the copies of the gene are placed in and the NGS analyser builds the DNA for every one of them. The general idea is that if it reads an adenine for example, at a certain position in 99% of the copies then we can be fairly certain that there is actually an adenine in this position in our own sample. By doing this over and over again, we ultimately end up knowing the sequence of the gene we extracted.
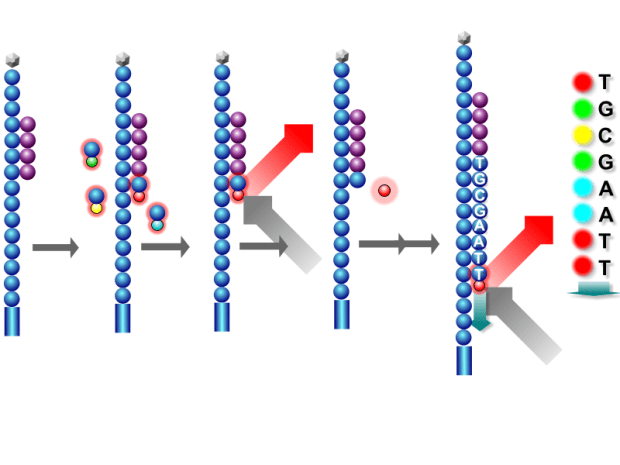 Ok, How Does That Actually Help?
Ok, How Does That Actually Help?
That all sounds great, but what can we do with this information?
The meat of the topic is of course the reasons why this sort of investigation is a useful tool. With these two techniques at play, we can detect mutations in a gene. We know that various genetic mutations cause various diseases so diagnostically speaking these techniques are important in confirming or ruling out diagnoses. Especially when it comes to certain types of cancer where the clinical symptoms are identical and the only differences are which mutations a patient may harbour. Secondly, if a patient has a previously unidentified disease, we can work backwards and find out if there are any underlying molecular changes that may be causing it. Insertions or deletion, copy number variations, translocations etc. can all be identified with these approaches. Thirdly, we can see how various drugs affect the body. For example, some drugs cause certain genes to be either over or under active and in some cases, either turned completely on or off! Having this wealth of information is invaluable to clinicians and scientists and is going to enable us to tackle human disease with a much greater understanding of the molecular abnormalities at the basis of these conditions.
Horizons
The future of molecular biology is fairly clear at this stage. Precision medicine. When it comes to sequencing the very genes which provide instructions to how the body works, we have unlocked a vast fountain of knowledge that was previously untapped. As part of this, scientists have been working for years to uncover why some treatments work for some and not other. The general idea that a “one size fits all” approach to medicine isn’t ideal, has a good basis. We know that people metabolise drugs differently, we know that they respond differently, and we know that by sequencing an individual’s genes we can find out what the best course of treatment is for that individual. Personalised medicine if you will. Although costs are still high for this type of science, as we get better and costs are reduced, you can expect to see it implemented on the frontline of medicine.
